In the previous post, I have shown how to use letsR to analyze species traits at the community level. But, in many cases this type of analysis can lead to spurious patterns (click here for further discussion on this issue). An alternative can be analyzing the data at the species-level. In this post, I will show two examples on how to make macroecological analysis at the species level using the letsR package. In the first example, we will continue the test of Rapoport’s rule on Phyllomedusa frogs using species centroids. In the second example, we will summarize climate spatial data at the species level to explore how temperature correlates with Phyllomedusa species extinction risk.
To start this test we can load our PresenceAbsence object that we generated previously (see the first and second posts on how to do it).
Note: I recommend to use the latest version of the letsR package on GitHub
library(letsR)data("PAM")To make things even more interesting, lets make an interactive map using the leaflet package (if you just want a normal plot use: plot(PAM)).
library(leaflet)
library(htmltools)r <- PAM$Richness_Raster
values(r)[values(r) == 0] <- NA
pal <- colorNumeric(c('#edf8e9','#c7e9c0','#a1d99b',
'#74c476','#31a354','#006d2c'),
values(r),
na.color = "transparent")
h = "https://mts1.google.com/vt/lyrs=s&hl=en&src=app&x={x}&y={y}&z={z}&s=G"
leaflet() %>%
addTiles(urlTemplate = h,
attribution = 'Google') %>%
addRasterImage(r, colors = pal, opacity = .6) %>%
addLegend(pal = pal, values = values(r),
title = "Species Richness")Example 1: Species level test of Rapoport’s rule on Phyllomedusa frogs.
Like the previous test at the community level, we first have to calculate species range size. We can do it directly on the species shapefiles for higher precision.
data("Phyllomedusa")
rangesize <- lets.rangesize(Phyllomedusa,
coordinates = "geographic")
rangesize <- rangesize / 1000 # Transform in km2The second step is to calculate species geographical centroid/midpoint using the function lets.midpoint. There are several ways to calculate species geographic centroid, and this function offers several methods to do it. When species range are both circular and continuous, all of the methods will provide the same answer. However, as the shape of distributions start to become more complex, different methods will provide very different answers. For this example, we will use the default option “PC” (polygon centroid). This method will generate a polygon from the raster, and calculate the centroid of this polygon.
centroids <- lets.midpoint(PAM)centroids| Species | x | y |
|---|---|---|
| Phyllomedusa araguari | -47.50000 | -19.500000 |
| Phyllomedusa atelopoides | -72.48928 | -7.090112 |
| Phyllomedusa ayeaye | -46.83259 | -20.836399 |
| Phyllomedusa azurea | -56.60321 | -19.316356 |
| Phyllomedusa bahiana | -40.16538 | -11.673186 |
| Phyllomedusa baltea | -74.50000 | -9.500000 |
| Phyllomedusa bicolor | -60.96245 | -3.443338 |
| Phyllomedusa boliviana | -62.00709 | -15.190434 |
| Phyllomedusa burmeisteri | -43.26027 | -18.045139 |
| Phyllomedusa camba | -66.00666 | -12.005590 |
| Phyllomedusa centralis | -55.50000 | -15.500000 |
| Phyllomedusa coelestis | -76.41726 | -4.587097 |
| Phyllomedusa distincta | -48.25991 | -25.652435 |
| Phyllomedusa duellmani | -77.50000 | -5.500000 |
| Phyllomedusa ecuatoriana | -78.50000 | -2.500000 |
| Phyllomedusa hypochondrialis | -55.66230 | -9.675312 |
| Phyllomedusa iheringii | -53.55319 | -31.863794 |
| Phyllomedusa itacolomi | -43.50000 | -20.500000 |
| Phyllomedusa megacephala | -43.00000 | -19.500773 |
| Phyllomedusa neildi | -69.50000 | 11.500000 |
| Phyllomedusa nordestina | -40.63072 | -10.860987 |
| Phyllomedusa oreades | -47.83385 | -15.168817 |
| Phyllomedusa palliata | -69.81883 | -9.452030 |
| Phyllomedusa perinesos | -77.50000 | -0.500019 |
| Phyllomedusa rohdei | -44.16199 | -22.126262 |
| Phyllomedusa sauvagii | -60.03017 | -24.154080 |
| Phyllomedusa tarsius | -67.10458 | -4.942237 |
| Phyllomedusa tetraploidea | -51.82583 | -24.139091 |
| Phyllomedusa tomopterna | -62.64377 | -4.179290 |
| Phyllomedusa trinitatis | -66.16788 | 10.368902 |
| Phyllomedusa vaillantii | -61.80198 | -5.069758 |
| Phyllomedusa venusta | -75.40027 | 7.667742 |
We can plot the geographical centroids using our cool leaflet maps. Dot size varies with range size in a log scale. You can pass the mouse over the dots to see the species name.
d <- data.frame("Species" = centroids[, 1],
"Range size" = rangesize)
sp <- sp::SpatialPointsDataFrame(centroids[, 2:3],
d)
h = "https://mts1.google.com/vt/lyrs=s&hl=en&src=app&x={x}&y={y}&z={z}&s=G"
leaflet(sp) %>%
addTiles(urlTemplate = h,
attribution = 'Google') %>%
addCircleMarkers(
col = rep("red", length(sp)),
stroke = F, fillOpacity = 0.6,
radius = log(rangesize)/4,
label = ~htmlEscape(Species),
labelOptions = labelOptions(noHide = F,
style = list(
"font-style" = "italic",
"font-size" = "14px",
"border-color" = "rgba(0,0,0,0.5)"
)))To check the Rapoport’s rule we can plot the latitude against the range size:
library(ggplot2)data_plot <- data.frame(centroids[, 2:3], "Range size" = rangesize)
g <- ggplot(data_plot, aes(x, Range_size))
g + geom_smooth() + geom_point() + labs(x = "Latitude(x)", y = "Range size")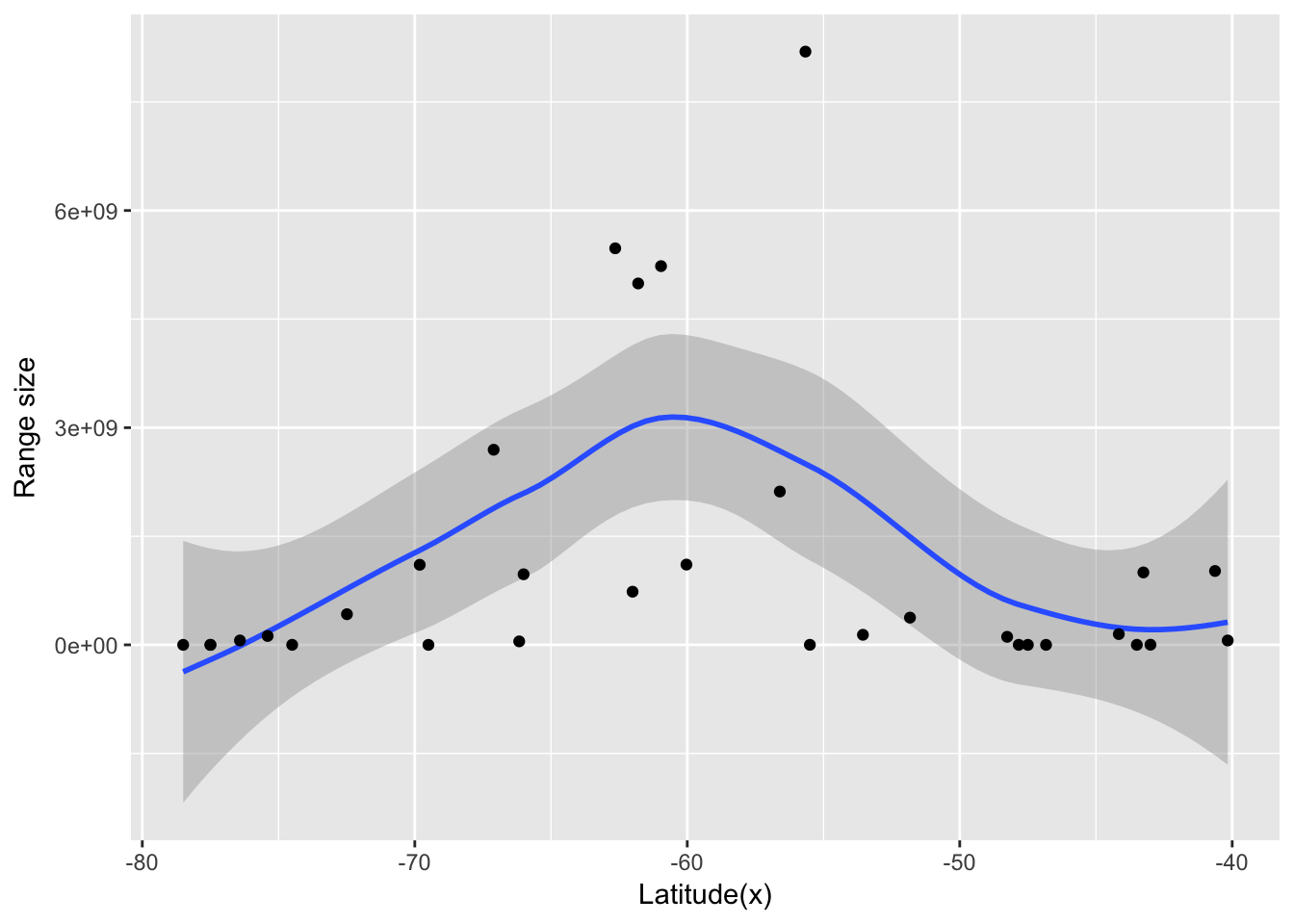
Again, the data indicate that Rapoport’s rule does not apply for Phyllomedusa genus. However, there seems to be an interesting pattern where range size decreases from the center towards the extremes of the group. This could be an effect of niche conservatism, where species in the extreme latitude would face very different conditions from the ancestral Phylllomedusa. Another possibility is that this pattern could be due to the shape of the continent, where extreme latitudes means smaller longitudes.
Example 2: Extinction risk correlation with temperature
To evaluate how temperature correlates with extinction risk, we first have to add the temperature variable to the PresenceAbsence object (click here for a detailed tutorial on how to add variables to a PresenceAbsence object).
r <- raster(getData("worldclim", var = "bio",
res = 10), 1) / 10
projection(PAM$Richness_Raster) <- projection(r)
PAM_env <- lets.addvar(PAM, r, fun = mean)Next step is to get the average temperature values per species. The lets.summarizer can do this directly on the resulting object of lets.addvar function (note that this can only be done if onlyvar = FALSE). We only have to indicate the position of the variable in the matrix using the argument pos.
pos <- which(colnames(PAM_env) == "bio1_mean")
temp_mean <- lets.summarizer(PAM_env, pos)temp_mean| Species | bio1_mean |
|---|---|
| Phyllomedusa araguari | 20.80840 |
| Phyllomedusa atelopoides | 25.70186 |
| Phyllomedusa ayeaye | 20.30772 |
| Phyllomedusa azurea | 23.50477 |
| Phyllomedusa bahiana | 22.99238 |
| Phyllomedusa baltea | 25.49704 |
| Phyllomedusa bicolor | 25.98631 |
| Phyllomedusa boliviana | 23.68508 |
| Phyllomedusa burmeisteri | 21.98282 |
| Phyllomedusa camba | 24.76891 |
| Phyllomedusa centralis | 23.89120 |
| Phyllomedusa coelestis | 24.68128 |
| Phyllomedusa distincta | 18.99102 |
| Phyllomedusa duellmani | 21.48040 |
| Phyllomedusa ecuatoriana | 15.60320 |
| Phyllomedusa hypochondrialis | 24.93516 |
| Phyllomedusa iheringii | 17.84105 |
| Phyllomedusa itacolomi | 19.76412 |
| Phyllomedusa megacephala | 21.19124 |
| Phyllomedusa neildi | 26.28497 |
| Phyllomedusa nordestina | 24.03002 |
| Phyllomedusa oreades | 23.70687 |
| Phyllomedusa palliata | 25.18994 |
| Phyllomedusa perinesos | 21.58766 |
| Phyllomedusa rohdei | 20.64380 |
| Phyllomedusa sauvagii | 22.31562 |
| Phyllomedusa tarsius | 25.46380 |
| Phyllomedusa tetraploidea | 20.46091 |
| Phyllomedusa tomopterna | 25.73224 |
| Phyllomedusa trinitatis | 25.43068 |
| Phyllomedusa vaillantii | 25.81837 |
| Phyllomedusa venusta | 24.54523 |
Following our example, we need to obtain the IUCN extinction risk data. Previous version of the package included functions to do this, but they are no longer supported. Luckily, there is a new package called rredlist that can do this for us. Yet, for now, we can use the example data in the letsR package called IUCN.
data("IUCN")IUCN| Species | Family | Status | Criteria | Population | Description_Year | Country |
|---|---|---|---|---|---|---|
| Phyllomedusa araguari | HYLIDAE | DD | Unknown | 2007 | Brazil | |
| Phyllomedusa atelopoides | HYLIDAE | LC | Unknown | 1988 | Bolivia, Brazil, Peru | |
| Phyllomedusa ayeaye | HYLIDAE | CR | B1ab(iii)+2ab(iii) | Unknown | 1966 | Brazil |
| Phyllomedusa azurea | HYLIDAE | DD | Unknown | 1862 | Argentina, Bolivia, Brazil, Paraguay | |
| Phyllomedusa bahiana | HYLIDAE | DD | Unknown | 1925 | Brazil | |
| Phyllomedusa baltea | HYLIDAE | EN | B1ab(iii)+2ab(iii) | Decreasing | 1979 | Peru |
| Phyllomedusa bicolor | HYLIDAE | LC | Stable | 1772 | Bolivia, Brazil, Colombia, French Guiana, Guyana, Peru, Suriname, Venezuela | |
| Phyllomedusa boliviana | HYLIDAE | LC | Stable | 1902 | Argentina, Bolivia, Brazil | |
| Phyllomedusa burmeisteri | HYLIDAE | LC | Stable | 1882 | Brazil | |
| Phyllomedusa camba | HYLIDAE | LC | Stable | 2000 | Bolivia, Brazil, Peru | |
| Phyllomedusa centralis | HYLIDAE | DD | Unknown | 1965 | Brazil | |
| Phyllomedusa coelestis | HYLIDAE | LC | Unknown | 1874 | Colombia, Ecuador, Peru | |
| Phyllomedusa distincta | HYLIDAE | LC | Decreasing | 1950 | Brazil | |
| Phyllomedusa duellmani | HYLIDAE | DD | Unknown | 1982 | Peru | |
| Phyllomedusa ecuatoriana | HYLIDAE | EN | B1ab(iii) | Decreasing | 1982 | Ecuador |
| Phyllomedusa hypochondrialis | HYLIDAE | LC | Stable | 1800 | Argentina, Bolivia, Brazil, Colombia, French Guiana, Guyana, Paraguay, Suriname, Venezuela | |
| Phyllomedusa iheringii | HYLIDAE | LC | Unknown | 1885 | Brazil, Uruguay | |
| Phyllomedusa itacolomi | HYLIDAE | DD | Unknown | 2006 | Brazil | |
| Phyllomedusa megacephala | HYLIDAE | DD | Unknown | 1926 | Brazil | |
| Phyllomedusa neildi | HYLIDAE | DD | Unknown | 2006 | Venezuela | |
| Phyllomedusa nordestina | HYLIDAE | DD | Unknown | 2006 | Brazil | |
| Phyllomedusa oreades | HYLIDAE | DD | Unknown | 2002 | Brazil | |
| Phyllomedusa palliata | HYLIDAE | LC | Stable | 1873 | Bolivia, Brazil, Ecuador, Peru | |
| Phyllomedusa perinesos | HYLIDAE | DD | Unknown | 1973 | Colombia, Ecuador | |
| Phyllomedusa rohdei | HYLIDAE | LC | Stable | 1926 | Brazil | |
| Phyllomedusa sauvagii | HYLIDAE | LC | Stable | 1882 | Argentina, Bolivia, Brazil, Paraguay | |
| Phyllomedusa tarsius | HYLIDAE | LC | Stable | 1868 | Brazil, Colombia, Ecuador, Peru, Venezuela | |
| Phyllomedusa tetraploidea | HYLIDAE | LC | Stable | 1992 | Argentina, Brazil, Paraguay | |
| Phyllomedusa tomopterna | HYLIDAE | LC | Stable | 1868 | Bolivia, Brazil, Colombia, Ecuador, French Guiana, Guyana, Peru, Suriname, Venezuela | |
| Phyllomedusa trinitatis | HYLIDAE | LC | Stable | 1926 | Trinidad and Tobago, Venezuela | |
| Phyllomedusa vaillantii | HYLIDAE | LC | Stable | 1882 | Bolivia, Brazil, Colombia, Ecuador, French Guiana, Guyana, Peru, Suriname, Venezuela | |
| Phyllomedusa venusta | HYLIDAE | LC | Decreasing | 1967 | Colombia, Panama |
Finally, we can verify the relationship between temperature and extinction risk.
level_order <- c("DD", "LC", "EN", "CR") # ordering for the plot
data <- data.frame("Status" = factor(IUCN$Status, levels = level_order),
"Temperature" = temp_mean[, 2])
g <- ggplot(data, aes(Status, Temperature))
g + geom_boxplot() + coord_flip()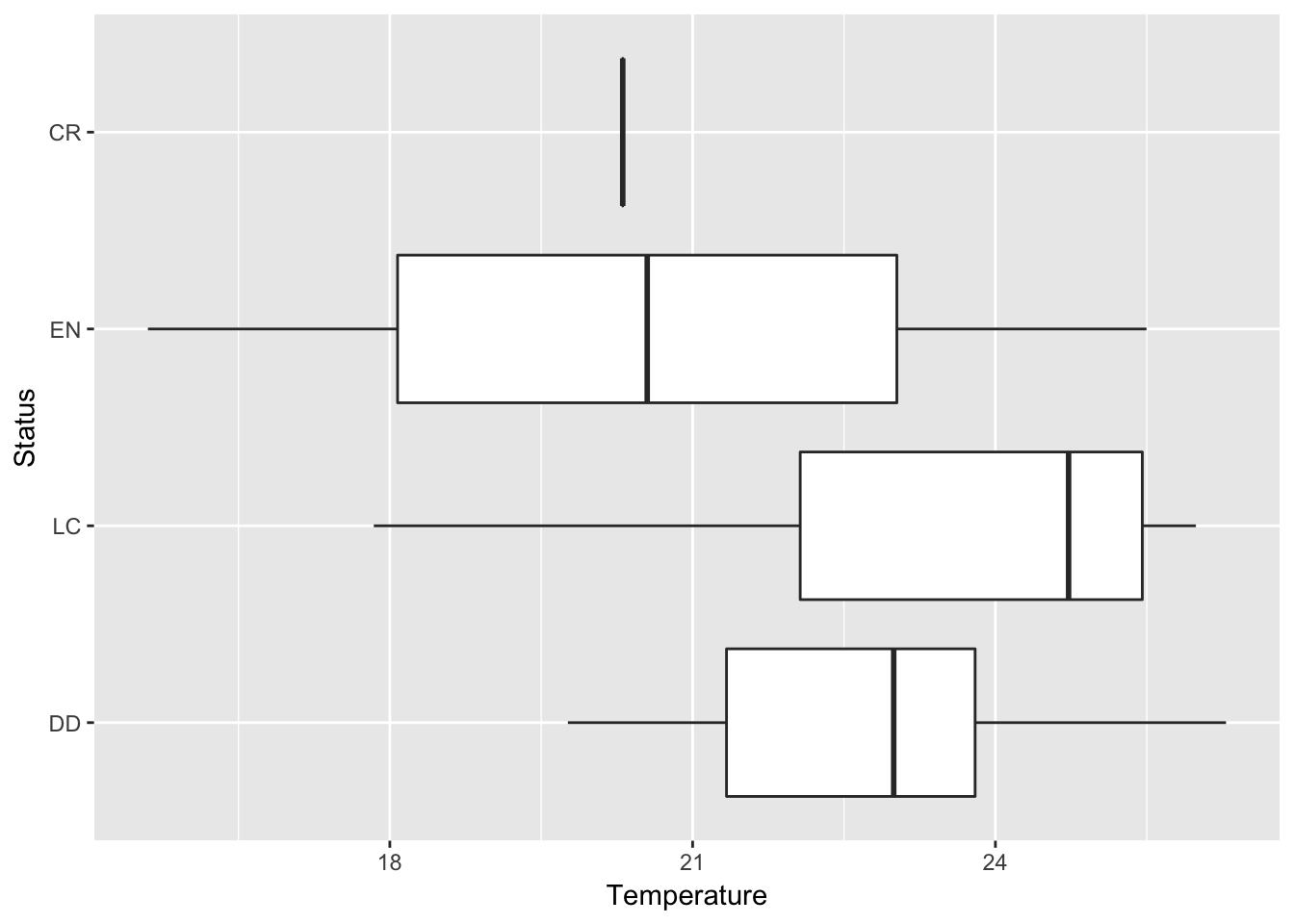
The graph indicate that there is a tendency for threatened Phyllomedusa species to occur in colder regions. Still, further statistical analysis should be done to confirm this pattern.
To cite letsR in publications use: Bruno Vilela and Fabricio Villalobos (2015). letsR: a new R package for data handling and analysis in macroecology. Methods in Ecology and Evolution. DOI: 10.1111/2041-210X.12401